 https://ibmm.umontpellier.fr/wp-content/uploads/Jao_Tanepau.jpg
250
200
Caroline Clavel
https://ibmm2.umontpellier.fr/wp-content/uploads/logo_ibmm_hd.jpg
Caroline Clavel2025-07-11 10:17:062025-07-11 10:17:06Congratulations to Joao for winning the Best Poster Award!
https://ibmm.umontpellier.fr/wp-content/uploads/Jao_Tanepau.jpg
250
200
Caroline Clavel
https://ibmm2.umontpellier.fr/wp-content/uploads/logo_ibmm_hd.jpg
Caroline Clavel2025-07-11 10:17:062025-07-11 10:17:06Congratulations to Joao for winning the Best Poster Award!- Read more
The Institut des Biomolécules Max Mousseron (IBMM, UMR 5247) is a research unit of UM, CNRS and ENSCM that develops research at the interface of chemistry, biology and health, integrating the challenges of sustainable development: feed, cure, protect.

News
 https://ibmm.umontpellier.fr/wp-content/uploads/Jao_Tanepau.jpg
250
200
Caroline Clavel
https://ibmm2.umontpellier.fr/wp-content/uploads/logo_ibmm_hd.jpg
Caroline Clavel2025-07-11 10:17:062025-07-11 10:17:06Congratulations to Joao for winning the Best Poster Award!
https://ibmm.umontpellier.fr/wp-content/uploads/Jao_Tanepau.jpg
250
200
Caroline Clavel
https://ibmm2.umontpellier.fr/wp-content/uploads/logo_ibmm_hd.jpg
Caroline Clavel2025-07-11 10:17:062025-07-11 10:17:06Congratulations to Joao for winning the Best Poster Award!
PhD offer: Revisiting the Amidation Reaction by Flow Mechanochemistry and Machine Learning (copie)
Context and objectives:
Mechanochemistry and flow chemistry…
16 research teams
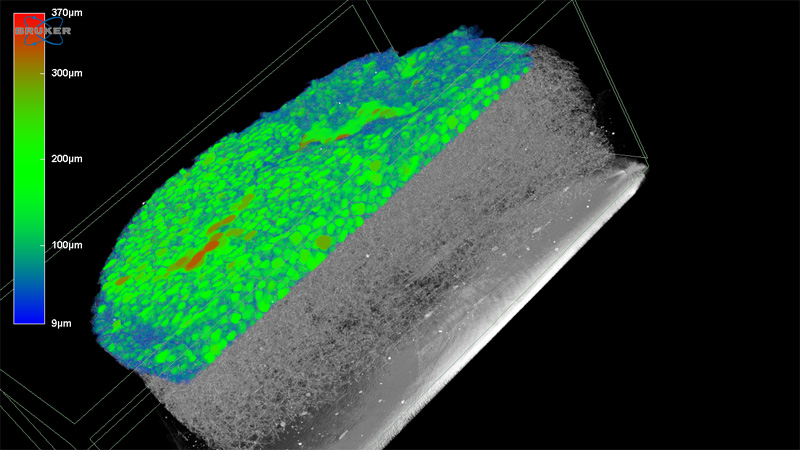

Polymers for Health and Biomaterials
30 March 2024/by Stephanie HERNANDEZ
Dynamics of Complex Biomolecular Systems
30 March 2024/by Stephanie HERNANDEZ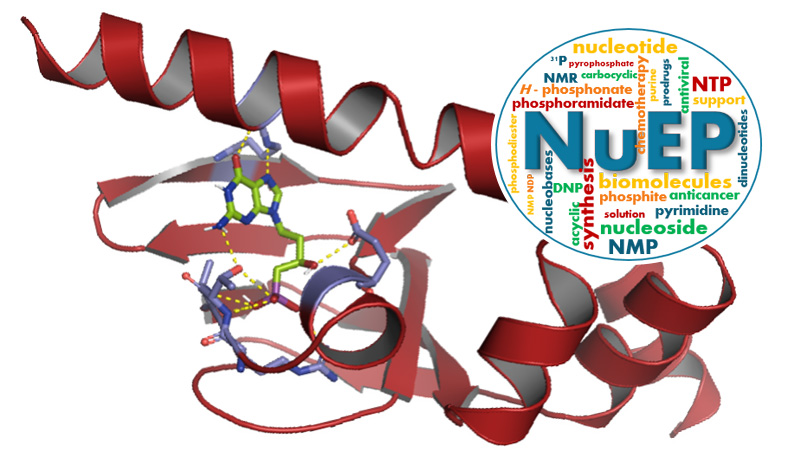
Nucleosides & Phosphorylated Effectors
30 March 2024/by Stephanie HERNANDEZ
ChemBioNAC
30 March 2024/by Stephanie HERNANDEZ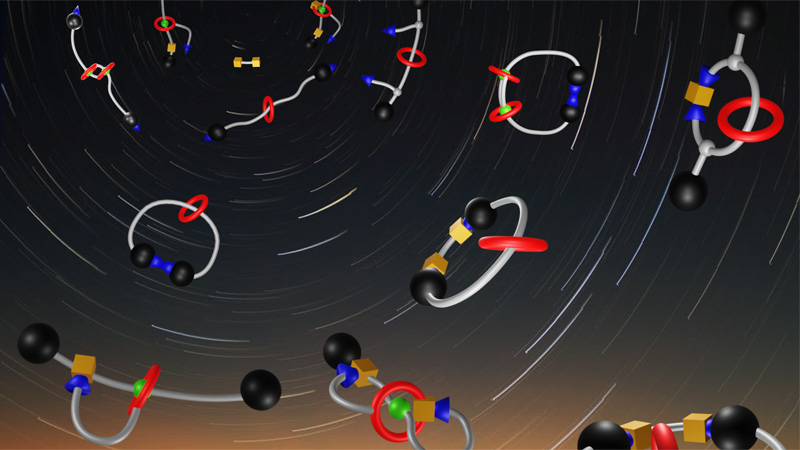
Supramolecular MAchines and ARchitecture Team
10 April 2024/by Stephanie HERNANDEZ
Oncotherapy and Oncopharmacology
19 December 2024/by Caroline Clavel
Green Chemistry and Enabling Technologies
30 March 2024/by Stephanie HERNANDEZ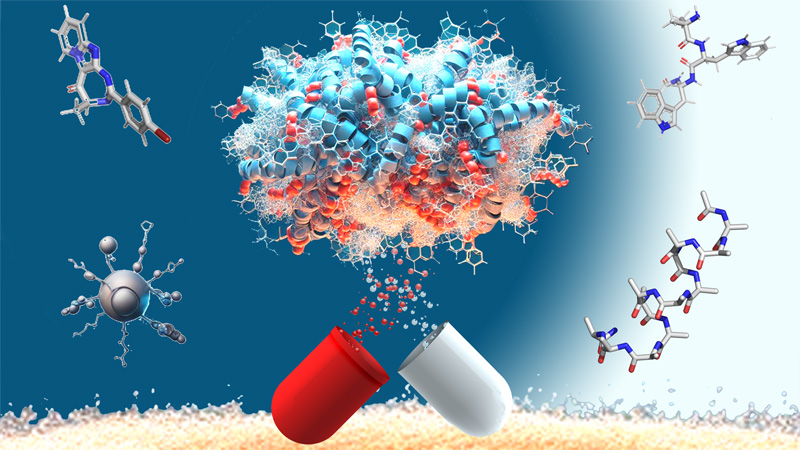
Amino acids, heterocycles, peptides & proteins
30 March 2024/by Stephanie HERNANDEZ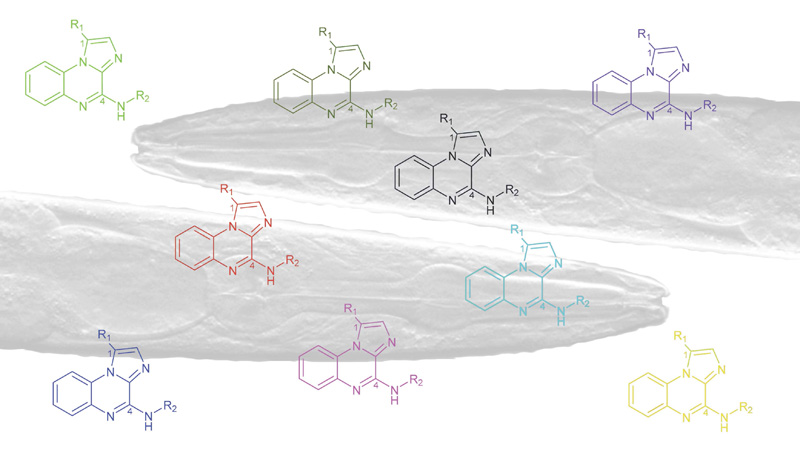
Oncopharmacochemy & Skin Pharmacotoxicology
6 April 2024/by Stephanie HERNANDEZ
Stereoselective Synthesis & Modified Amino Acids
30 March 2024/by Stephanie HERNANDEZ
Glycochemistry & molecular recognition
30 March 2024/by Stephanie HERNANDEZ
Analytical Sciences of Biomolecules
30 March 2024/by Stephanie HERNANDEZ
Cellular pharmacology
30 March 2024/by Stephanie HERNANDEZ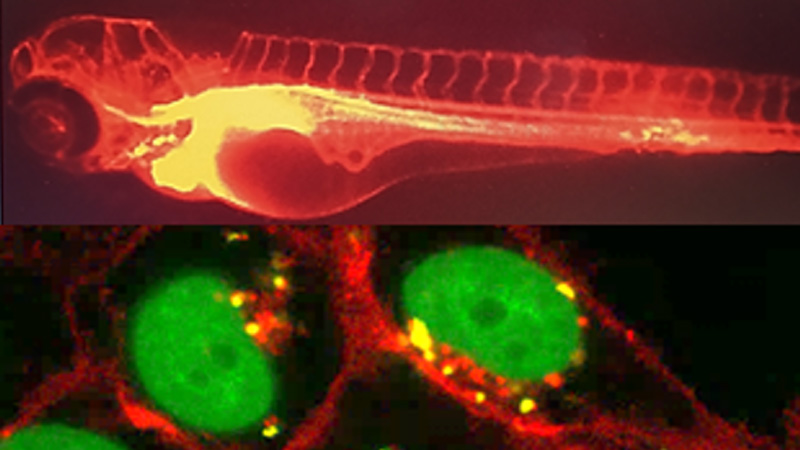
Glyco and nanovectors for therapeutic targeting
30 March 2024/by Stephanie HERNANDEZ
Pharmacochemistry Synaptic Transmission & Neuroprotection
30 March 2024/by Stephanie HERNANDEZ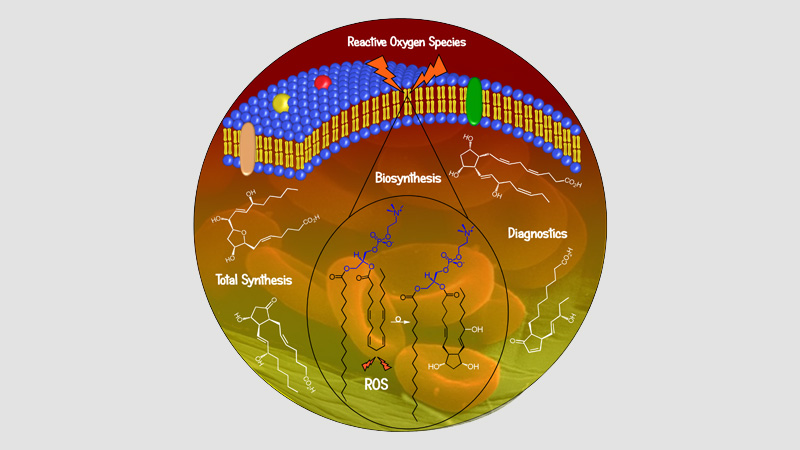







PhD offer: PHOTOCLICK: Development of biorthogonal photo-crosslinking reactions for 3D printing micro-structured biomaterials
Contract: 3 years starting from October 2025 Application deadline: 04/07/2025 Location: Institute of Biomolecules Max Mousseron (IBMM), Montpellier, France and Institute for Molecular Systems Engineering and Advanced Materials (IMSEAM), Heidelberg, Germany The project This thesis project is located at the interface of click chemistry, photochemistry, materials, 3D printing and biology. The goal is to […]
PhD offer: Filled !
ADVANCED 4D BIOMATERIALS FOR MUCOSA AND SUB-MUCOSA TREATMENT IN PATIENTS AFFECTED BY INTESTINAL DISEASES (Daedalus) financed by the EU program HORIZON EUROPE 4D biomaterials enable new surgical treatments by autonomously acting in response to environmental stimuli, thus overcoming the limitations of standard medical strategies. DAEDALUS is a European project that gathers 13 partners and […]