Home > Research > PeptiSystems
ANR Project PeptiSystems (2014–2019)
Energetic processes driving potential peptide protometabolisms at the origin of living systems
PeptiSystems is a collaborative research project on Prebiotic Chemistry and Origins of Life between DSBC (Montpellier) and PIIM (Marseille) groups, granted by the Agence Nationale de la Recherche (ANR) for four years, project number ANR-14-CE33-0020.
Project outline
The ongoing detection of numerous exoplanets with conditions becoming more and more similar to those of the Earth, places the question of the plurality of living worlds at the forefront of unresolved scientific issues. However, studies on the possible nature and emergence of other forms of life, as well as their detection can only be undertaken in an interdisciplinary way. Life on Earth constitutes a historical process that developed by complexification starting from chemical systems. In the PeptiSystems project we consider that both physico-chemical driving forces and contingency are responsible for this process since its early beginning, which means that non-historical aspects deserve a scientific approach, which is of general interest in astrobiology.
The driving force towards complexity
The theoretical developments will constitute the starting point of experimental studies of this project and a matter of ongoing investigations to contribute to a better understanding of the physico-chemical origin of complexity. Within this perspective, the emergence of protometabolisms defined as networks of chemical reactions proceeding under far-from-equilibrium conditions and involving nonlinear features and therefore being capable of generating self-organisation (associated with a local decrease in entropy compensated by the irreversibility of the overall process), is a key step for the origin of life.
Irreversible reaction networks
Chemical networks of this kind must work as unidirectional (kinetically irreversible) sequences of reactions or preferably as unidirectional reaction cycles in which a further process of positive chemical feedback would be capable of generating systems endowed with autocatalytic properties and then behaving in a nonlinear way. Determining which pathways could have constantly or repeatedly fed these systems with energy to maintain the far from equilibrium state is then essential to understand how self-organisation could emerge.
Biopolymers
This approach is applied to the formation of biopolymers capable of functional activities that is generally considered as a prerequisite for the development of life. Our original goal is to propose an overall scenario integrating the formation of peptides and other related processes to build a network capable of giving rise to emergent properties related to the connections of the different parts of the network.
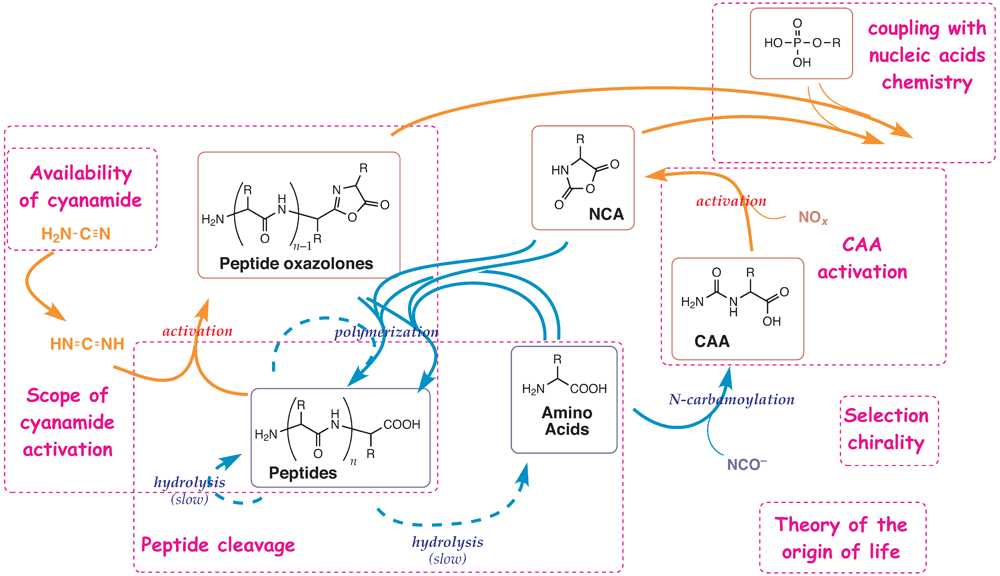
The chemical network constituting a peptide protometabolism
to be investigated more deeply in the project
Peptide formation and translation
The PeptiSystems project is aimed at (i) understanding how peptides could be formed under prebiotic conditions and how energy sources could have been coupled to peptide bond formation. But it is not limited to this goal since peptides made from racemic mixtures of amino acids are unlikely to adopt definite structures needed for specific activity, so that we will (ii) address the question of the emergence of selectivity (stereoselectivity and sequences with single handedness or defined structure) or that of improbable but dynamically stable states.
Lastly, (iii) the emergence of translation at an early stage of evolution suggests that the chemistry of amino acids and peptides can be coupled to that of an information carrier supporting the hypothesis of a peptide-nucleotide co-evolution. Developing knowledge about chemical pathways through which amino acids or short peptides could have interacted with ribonucleotides monomers and oligomers is a prerequisite for understanding the emergence of the translation process and will constitute a priority in our investigations.
All these topics are experimentally investigated by monitoring relevant reactions through usual methods of organic and analytical chemistry (HPLC, NMR, MS, UV spectrometries…).
People involved in ANR framework:
DSBC group of IBMM (Montpellier)
Permanent staff:
Robert Pascal (CNRS emeritus Senior Scientist, project leader) [appointed at PIIM/ASTRO from 2019-05-01]
Laurent Boiteau (CNRS Associate Scientist)
Jean-Christophe Rossi (Associate Professor, University of Montpellier)
Former staff :
Damien Beaufils (MESR graduate student, promoted 06 Nov. 2015)
Ziwei Liu (Simons Foundation postdoctoral fellow, nov. 2013 – jan 2017)
Ghinwa Ajram (graduate student / ANR — promoted 29 Nov. 2018)
ASTRO group of PIIM lab (Marseille) [website]
Permanent staff:
Grégoire Danger (Full Professor, Aix-Marseille University)
Fabrice Duvernay (Associate Professor, Aix-Marseille University)
External collaborations (not sponsored by ANR)
Laboratory SQPOV, UMR 408 INRA, UAPV Université d‘Avignon et des Pays de Vaucluse, France
– Dr. Raphaël Plasson
Ben Gurion University of Negev, Israel
– Prof. Addy Pross [website]
MRC Laboratory of Molecular Biology, Cambridge, UK
[website]
– Prof. John D. Sutherland (group leader),
– Dr. Ziwei Liu (postdoctoral fellow since March 2017),
– Dr. Dougal Ritson (career development fellow)
IAS (Information and Autonomous Systems) Research Centre for Life, Mind & Society,
EHU-UPV University of Basque Country, Spain
[website]
– Prof. Kepa Ruiz-Mirazo (group leader),
– Sara Murillo Sánchez (graduate student, promoted 07 Sept. 2017)
UCSB University of Califdornia Santa Barbara & NASA, USA
– Prof. Irene A. Chen
Laboratory ICBMS, UMR 5246 CNRS, Université Claude-Bernard Lyon 1, France
– Prof. Pierre Strazewski (group leader),
– Dr. Michele Fiore (Maître de Conférences)
Publications
(“Open Acess” articles have a DOI link labelled with an icon )
R. Pascal,
Kinetic barriers and self-organisation of life.
Isr. J. Chem. 2015, 55, 865–874.
DOI: 10.1002/ijch.201400193
R. Pascal, A. Pross,
Stability and its manifestation in the chemical and biological worlds.
Chem. Commun. 2015, 51(90), 16160–16165.
DOI: 10.1039/C5CC06260H
D. Beaufils, S. Jepaul, Z. Liu, L. Boiteau, R. Pascal,
The activation of free dipeptides promoted by strong activating agents in water does not yield diketopiperazines.
Orig. Life Evol. Biosph. 2016, 46(1), 19–30.
DOI: 10.1007/s11084-015-9455-0
S. Murillo Sánchez, D. Beaufils, J. M. González Mañas, R. Pascal, K. Ruiz-Mirazo,
Fatty acids’ double role in the prebiotic formation of a hydrophobic dipeptide.
Chem. Sci. 2016, 7(5), 3406–3413.
DOI: 10.1039/c5sc04796j
R. Pascal,
Physicochemical requirements inferred for chemical self-organization hardly support an emergence of life in the deep ocean of icy moons.
Astrobiology 2016, 16, 328–334.
DOI: 10.1089/ast.2015.1412
 Z. Liu, L. Rigger, J.-C. Rossi, J. D. Sutherland, R. Pascal,
Z. Liu, L. Rigger, J.-C. Rossi, J. D. Sutherland, R. Pascal,
Mixed anhydride intermediates in the reaction of 5(4H)-oxazolones with phosphate esters and nucleotides.
Chem. Eur. J. 2016, 22(42), 14940–14949.
DOI: 10.1002/chem.201602697
□ Inside cover.
Chem. Eur. J. 2016, 22(42), 14758.
DOI: 10.1002/chem.201603523
R. Pascal, A. Pross,
The logic of life.
Orig. Life Evol. Biosph. 2016, 46(4), 507–513.
DOI: 10.1007/s11084-016-9494-1
R. Pascal, A. Pross,
A roadmap toward synthetic protolife.
Synlett 2017, 28(01), 30–35.
DOI: 10.1055/s-0036-1589403
Z. Liu, C. Hanson, G. Ajram, L. Boiteau, J.-C. Rossi, G. Danger, R. Pascal,
5(4H)-Oxazolones as effective aminoacylation reagents for the 3’-terminus of RNA.
Synlett 2017, 28(01), 73–77.
DOI: 10.1055/s-0036-1588647
A. Pross, R. Pascal,
How and why kinetics, thermodynamics, and chemistry induce the logic of biological evolution.
Beilstein J. Org. Chem. 2017, 13, 665–674.
DOI: 10.3762/bjoc.13.66
N. Abou Mrad, G. Ajram, J.-C. Rossi, L. Boiteau, F. Duvernay, R. Pascal, G. Danger,
The prebiotic C-terminal elongation of peptides can be initiated by N-carbamoyl amino acids.
Chem. Eur. J. 2017, 23(31), 7418–7421.
DOI: 10.1002/chem.201700702
Z. Liu, J.-C. Rossi, R. Pascal,
How prebiotic chemistry and early life chose phosphate.
Life 2019, 9(1), 26.
DOI: 10.3390/life9010026
Z. Liu, G. Ajram, J.-C. Rossi, R. Pascal,
The chemical likelihood of ribonucleotide-α-amino acid copolymers as players for early stages of evolution.
J. Mol. Evol. 2019, 87(2–3), 83–92.
DOI: 10.1007/s00239-019-9887-7
A. D. Pressman, Z. Liu, E. Janzen, C. Blanco, U. F. Müller, G. F. Joyce, R. Pascal, I. A. Chen,
Mapping a systematic ribozyme fitness landscape reveals a frustrated evolutionary network for self-aminoacylating RNA.
J. Am. Chem. Soc. 2019, 141(15), 6213–6223.
DOI: 10.1021/jacs.8b13298
R. Pascal,
A possible prebiotic basis for metabolism.
[invited News & Views commentary]
Nature 2019, 569, 47–49.
DOI: 10.1038/d41586-019-01322-3
R. Pascal, A. Pross,
Chemistry’s kinetic dimension and the physical basis for life.
J. Syst. Chem. 2019, 7, 1–8.
DOI n/c
G. Ajram, J.-C. Rossi, L. Boiteau, R. Pascal,
A reassessment of prebiotically relevant chemical agents for the activation of α-amino acids and peptides.
J. Syst. Chem. 2019, 7, 19–28.
DOI n/c
R. Pascal, I. A. Chen,
From soup to peptides.
[invited News & Views commentary]
Nat. Chem. 2019, 11, 763–764.
DOI: 10.1038/s41557-019-0318-6
Links
- Agence Nationale de la Recherche
- Physique des Interactions Ioniques et Moléculaires – UMR7345, Marseille
- Société Française d’Exobiologie (French Astrobiology Society)
- ISSOL – The International Astrobiology Society
- EU COST Chemistry Action CM1304 Emergence and Evolution of Complex Chemical Systems
- EU COST TransDomain Action TD1308 Life-Origins
- Simons Collaboration on the Origins of Life (Simons Foundation, USA)

The Montpellier PeptiSystems team in June 2014
(from left to right: Jean-Christophe, Sara, Laurent, Damien, Robert, Ziwei)